Using Comet to track your experiments can improve your performance as a Data Scientist because Comet helps you to organize your code, compare your implemented models, and select the best model. Once you have selected the best model, you can build a prediction service, which you can then send to production for implementation. And here is where DevOps comes in, with all its best practices and strategies to run your code.
DevOps permits you to integrate the development phase with the operational one, in which all the context around your code should be configured. The context around the code could include the environment variables, the required packages needed to run the code, and so on.
To integrate the software with its context, you may use a Docker image, which is a standalone runnable system, which you can run everywhere, provided that the hosting machine runs a Docker engine. In the previous example, you can build a Docker image that contains your prediction service, and then run it in a Docker container.
Many tutorials exist on how to build a Docker image to run your code. But what about building a Docker image for your Comet experiments? It would be great if you built a Docker image that tracks your experiments in Comet! You could download it from the Docker Hub, then configure it with your Comet API key, and finally run it in a Docker container.
In this article, I will describe exactly this process, by means of an example.
The article is organized as follows:
- Overview of the use case
- Wrapping the Comet experiment in a Docker image
Overview of the use case
As an example, I use the well-known diabetes dataset, provided by the scikit-learn library, and I build a Linear Regression model. Then, I calculate Mean Squared Error (MSE) and R2 score, and I log them as metrics in Comet.
Firstly, I import all the required libraries:
import os
from comet_ml import Experiment
from sklearn.datasets import load_diabetes
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error, r2_score
Secondly, I load the Comet configuration parameters from the environment variables. This will be useful when I will build the Docker image:
COMET_API_KEY = os.environ.get("COMET_API_KEY")
COMET_WORKSPACE = os.environ.get("COMET_WORKSPACE")
COMET_PROJECT = os.environ.get("COMET_PROJECT")
I load the dataset, and I split it into training and test set:
data = load_diabetes() X = data.data y = data.targetX_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
I build the Linear Regression model, and I calculate MSE and R2 score:
model = LinearRegression() model.fit(X_train, y_train) y_pred = model.predict(X_test)mse = mean_squared_error(y_test, y_pred) r2_score = r2_score(y_test, y_pred)
Finally, I build a Comet experiment, and I log the calculated metrics:
experiment = Experiment( api_key=COMET_API_KEY, project_name=COMET_PROJECT, workspace=COMET_WORKSPACE, )experiment.log_metric('mse', mse) experiment.log_metric('r2', r2_score)
I can test the code as follows. In a terminal, I set the Comet parameters:
export COMET_API_KEY=MY_API_KEY
export COMET_PROJECT=MY_COMET_PROJECT
export COMET_WORKSPACE=MY_COMET_WORKSPACE
and then I run the script:
python test_model.py
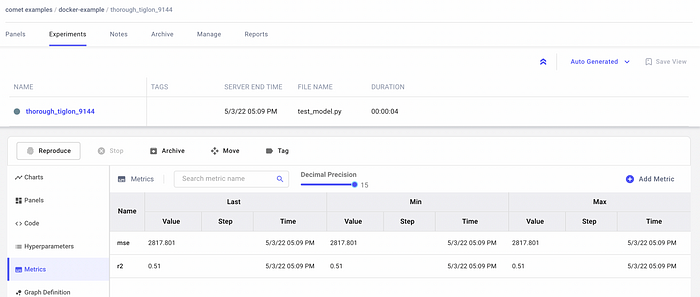
I should be able to see the results in Comet, as shown in the following figure:
Wrapping the Comet experiment in a Docker image
Before building a Docker image with my Comet experiment, I need to download and install Docker Desktop. Once installed, I can proceed as follows.
I need to write two files:
- requirements.txt
- Dockerfile
The requirements.txt file contains all the Python libraries required by the previous script. It is a plain text, that contains a library per line, as shown in the following piece of code:
comet_ml
scikit-learn
The Dockerfile contains the directives used to build the Docker image. It defines the base image from which you should build the image. In my case, I can use a slim version of Python:
FROM python:slim
Now, I specify that I want to copy all the content of the current directory to the image:
COPY . .
Next, I install all the required packages:
RUN pip install --no-cache-dir -r requirements.txt
Finally, I set the operation to perform when the image is run in a Docker container:
CMD ["python", "./test_model.py"]
I save the file as Dockerfile.
Eventually, I am ready to build the Docker image. I open a terminal, and I enter the directory containing all my files:
From that directory, I run the following command:
docker build -t cm .
The -t parameter specifies the name of the docker image. The building process starts.
When the process finishes, I should be able to see your image by running the following command:
docker images
which should give an output similar to the following one:

Before running the image in a Docker container, I need to specify the environment variables. I can define a file, named env.list, which contains the Comet configuration parameters:
COMET_API_KEY=MY_COMET_API
COMET_PROJECT=MY_COMET_PROJECT
COMET_WORKSPACE=MY_COMET_WORKSPACE
Finally, I can run the Docker image in a Docker container:
docker run --env-file /path/to/env.list --rm cm
You should be able to view the results in Comet!
Summary
Congratulations! You have just learned how to integrate a Comet experiment in a Docker image, and then run it as a Docker container! The process is quite simple, you only need to write:
- the code of your Comet experiment
- the
requirements.txtfile with all the required libraries - the
Dockerfilewith the building process.
Comet is very powerful to build and track your experiments. You may also be interested in other features provided by Comet, including How to Write your Comet Experiments in R and Writing a Classification Task using Comet and Java.
Stay tuned for new tutorials on Comet and don’t forget you can get started for free!
Happy Coding! Happy Comet!
